About the CoMSES Model Library more info
Our mission is to help computational modelers develop, document, and share their computational models in accordance with community standards and good open science and software engineering practices. Model authors can publish their model source code in the Computational Model Library with narrative documentation as well as metadata that supports open science and emerging norms that facilitate software citation, computational reproducibility / frictionless reuse, and interoperability. Model authors can also request private peer review of their computational models. Models that pass peer review receive a DOI once published.
All users of models published in the library must cite model authors when they use and benefit from their code.
Please check out our model publishing tutorial and feel free to contact us if you have any questions or concerns about publishing your model(s) in the Computational Model Library.
We also maintain a curated database of over 7500 publications of agent-based and individual based models with detailed metadata on availability of code and bibliometric information on the landscape of ABM/IBM publications that we welcome you to explore.
Displaying 2 of 2 results finite automata clear search
Peer reviewed Flibs’NFarol: Self-Organized Efficiency and Fairness Emergence in an Evolutive Game
Cosimo Leuci | Published Thursday, October 12, 2023According to the philosopher of science K. Popper “All life is problem solving”. Genetic algorithms aim to leverage Darwinian selection, a fundamental mechanism of biological evolution, so as to tackle various engineering challenges.
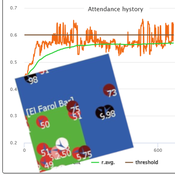
Flibs’NFarol is an Agent Based Model that embodies a genetic algorithm applied to the inherently ill-defined “El Farol Bar” problem. Within this context, a group of agents operates under bounded rationality conditions, giving rise to processes of self-organization involving, in the first place, efficiency in the exploitation of available resources. Over time, the attention of scholars has shifted to equity in resource distribution, as well. Nowadays, the problem is recognized as paradigmatic within studies of complex evolutionary systems.
Flibs’NFarol provides a platform to explore and evaluate factors influencing self-organized efficiency and fairness. The model represents agents as finite automata, known as “flibs,” and offers flexibility in modifying the number of internal flibs states, which directly affects their behaviour patterns and, ultimately, the diversity within populations and the complexity of the system.
Peer reviewed Flibs'NLogo - An elementary form of evolutionary cognition
Cosimo Leuci | Published Thursday, January 30, 2020Flibs’NLogo is an agent-based simulation implemented in NetLogo that models the evolution of perfect predictors through a genetic algorithm. The agents, called flibs (finite living blobs), are finite‑state automata whose behaviour is encoded in circular chromosomes. They inhabit a “primordial computer soup” and are tasked with anticipating a user‑defined periodic binary sequence. Each generation consists of 100 evaluation cycles, during which a flib’s fitness is incremented each time its output correctly matches the next environmental signal.
Reproduction follows an elitist scheme: a donor (current fittest individual) replaces a randomly chosen recipient either by cloning (complete genome substitution) or by bacterial‑like conjugation (unidirectional horizontal transfer of a random chromosome segment). A stochastic mutagenesis operator introduces point mutations in genes, while the reproductive strategy gene can also switch under a mixed-reproduction regime. Population dynamics are monitored via genomic diversity indices (Shannon‑Wiener, Simpson), a phenotypic simpleness metric that distinguishes the low number of states actually used from the genomic potential.
The model serves as a digital evolutionary laboratory for exploring the interplay among bounded rationality, collective adaptation, and the emergence of anticipatory behaviour. By linking evolutionary computation with cognitive concepts, Flibs’NLogo investigates fundamental transitions from reactive to predictive systems and allows for testing whether populations evolve toward minimal necessary complexity or exhibit an intrinsic drift toward structural elaboration.